R and RStudio Install: The User Experience, Part 2
Time to get your hands dirty...
Data Science Altitude for This Article: Base Camp.
Today’s post goes into the ‘meat and potatoes’ of the install process for R.
If you look to get more involved in a data science career or a more classic application development job in Java, .NET, or something similar, it’s important to develop a comfort level with installing software on your own machine. You’ll have to do it repeatedly over your career. Even if you work in a locked-down environment that delegates machine upkeep to other people and processes, if you are truly curious about your craft you’ll have a home machine with your own home-brew of software and projects.
All the best things in life happen when your curiosity engages you into action. So practice makes perfect, as they say. Let’s practice!
Installation of R - It’s All About the CRAN
When in doubt, go to the source. For R, that’s the front page for r-project.org. From there, you’ll notice a suggestion to download it from your nearest CRAN mirror. CRAN is short for ‘Comprehensive R Archive Network’ and consists of a set of servers worldwide that are dedicated to hosting current and past versions of the language.
For me, I could have picked either the CRAN mirror at Washington University in St. Louis or the one at the University of Kansas. Whichever one you pick will take you to a page where we now start to get down to business. Remember that choice later, you’ll have a setting where you need to specify a CRAN mirror to service ongoing updates.
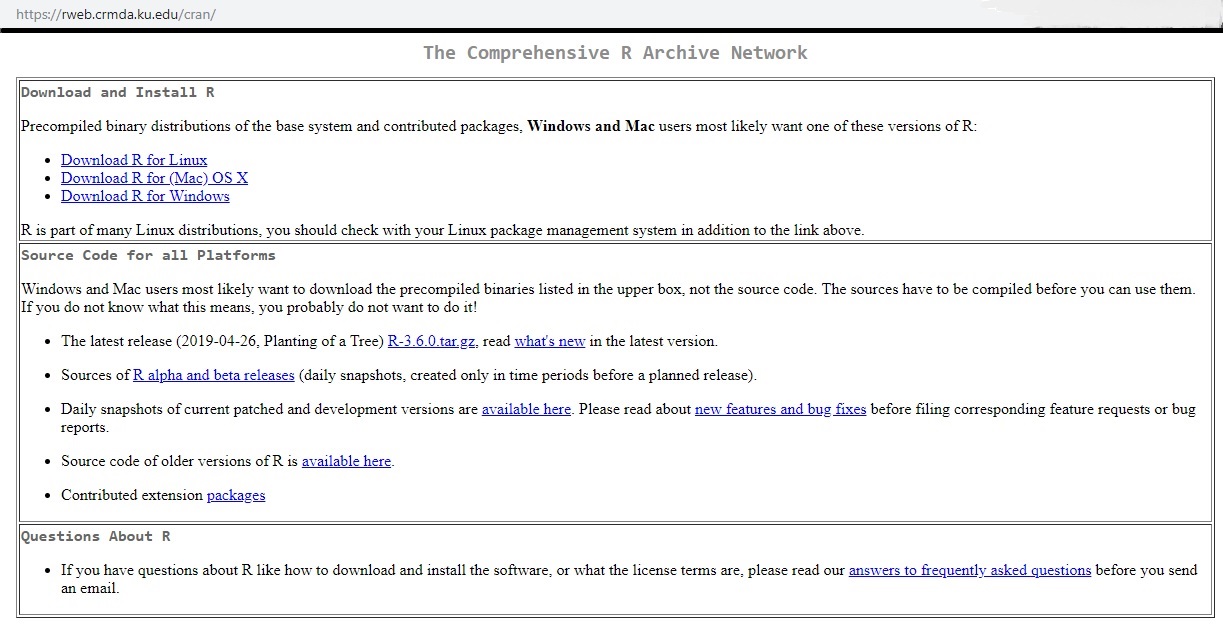
Once you choose whether you’re downloading for Linux, Mac, or Windows, you’re presented with another choice of platform to install on. I don’t have a Mac, and I’ll assume that if you have a Linux box or instead run a Linux virtualization on your machine (I use VirtualBox on my laptop for a virtual Linux/Ubuntu environment) that you won’t need much help from me. So I’ll proceed on the Windows side.
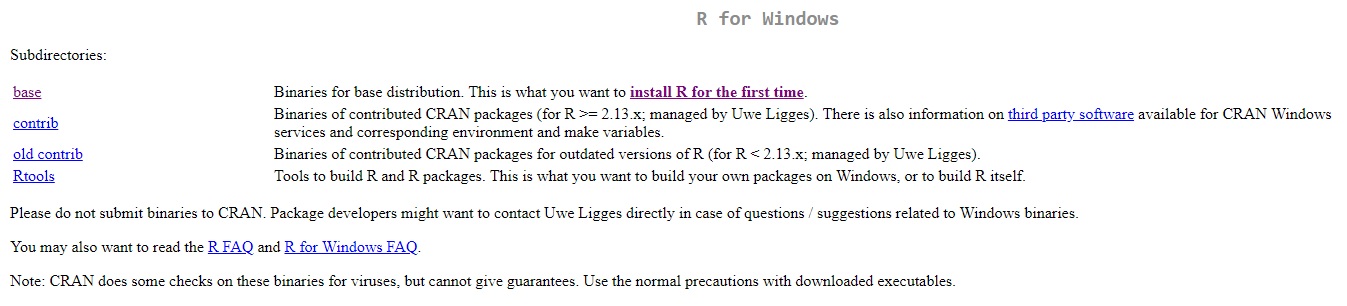
When you click on “Download R for Windows”, you’re presented with a list. At the top is the option to download the base distribution. While we won’t touch on it further today, note the presence of the Rtools link. For more advanced users it’s an assemblage of utilities for - among other things - building packages from their original source code.
Checking Out the FAQ
This takes you to the page where you can download the latest version. The compressed payload is only 80 MB so even those of you with bad throughput won’t notice. For the beginner, some of the questions here are only of passing interest if you’re not familiar with them. The reference to md5sum relates to a validation process by which one can tell if an object that was transferred to you is exactly the same as the one at the source. Ones and zeroes out of sequence can be a bad thing… Read up further on that if interested.
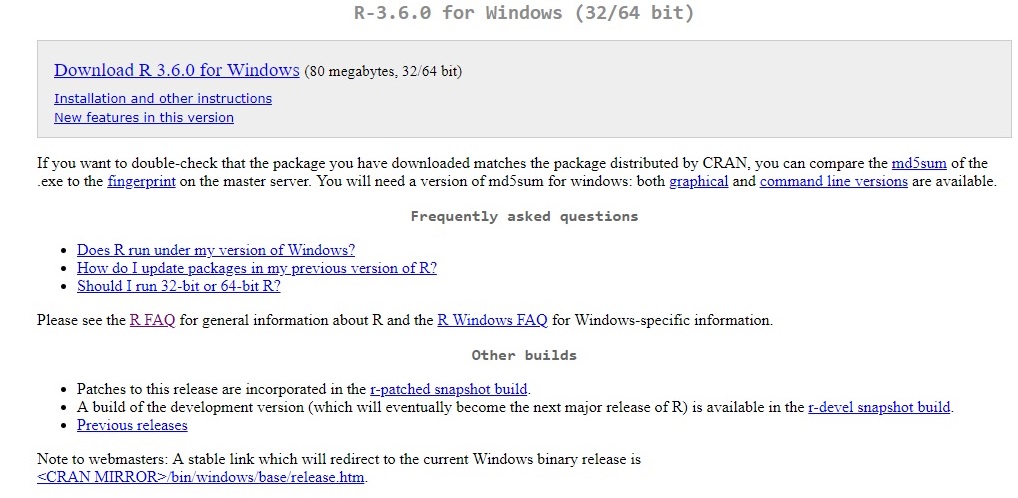
For the items in the ‘Frequently asked questions’ section, though, if you’re not sure what they’re talking about I’d read them. They’re subsections of the main R FAQ document. Long story short, unless you’re running a really old machine, you should be good for your operating system version and should be able to run the 64-bit version of R (or anything else):
Firing Up R for the First Time
Clicking on “Download R 3.6.0 for Windows” - takes us further along and pulls a R-3.6.0-win executable file (above) into your browser’s download folder. By the time you read this, another version might be the most recent. Once it’s downloaded, double-click on it to get it rolling.
Unless you want to put it on something other than your C: drive (if you have a dual-drive or otherwise-partitioned machine), take the defaults when asked by the installer. The only exception to that is whether you want to check the boxes that put a shortcut to run R natively on the desktop after the install. Your choice, but if we’re installing R Studio as our environment, it’s not necessary in my opinion. One more cluttering desktop shortcut…
I would, however, find the ‘R x64 3.6.0’ program and fire it up. Again, installing a version in the future will result in a different version number.
While it’s certainly possible to do our initial configuration of R in its graphical interface (like setting the default CRAN mirror for updates), I have historically done it within RStudio at the same time that I customize RStudio for the first time.
The following post is about the installation of RStudio and a brief overview of some of the cool things we can do with it.
Further Information on the Subject:
Another repository is available outside of CRAN for R packages and focuses on the needs of the biological and genomic research community. While not for the beginner, Bioconductor has packages that provide high-quality data analysis for publication in the likes of the Journal of the American Medicine Association (JAMA).
